The authors obtained de-identified genome-wide expression profiles from clinical use of the Decipher RP test in 9,953 men from the GRID registry database. The physician reported patient race was annotated according to US Census Bureau categories as one of Black/African-American, White (non-Hispanic), White (Hispanic), Asian or Other (American Indian, Native Alaskan or Hawaiian/Pacific Islander). Multivariable logistic regression odds ratio (OR) analysis was used to assess the association of 82 GRID signatures with African-American (AAM) as compared to non-African American (non-AAM) race status.
A subset of 1,043 men with known race status in the GRID including 529 African American (51%), 23 Asian (2%), 14 White (Hispanic) (1%), 472 White (Non-Hispanic) (45%) and 5 Other, were analyzed. The median age at surgery was 63 years (range, 41-81 years). AAM tended to be younger (p<0.004), had higher median preoperative PSA (p<0.001), lower Grade Group disease (p=0.04), higher rates of positive margins (p<0.001) and bladder neck invasion (p=0.01) as compared to non-AAM men. Decipher risk categories were not significantly different between the groups (p0.6). Non-AAM men had more ERG (40% vs 23%, p<0.001) and ETS (10% vs 6%, p=0.02) expression, whereas AAM had more Triple Negative (48% vs 39%, p<0.005) and SPINK1 (23% vs 11%, p<0.001) molecular subtype tumors (Figure 1). After adjusting for clinical and pathological risk features in multivariable analysis, AAM were less likely to be PAM50 Luminal (A or B, OR: 0.58, p=0.01), more Basal-like (OR: 2.1, p<0.001), had lower Androgen Receptor activity (OR: 2.1, p=0.01) and high PORTOS radiation response (OR: 2.2, p=0.01). AAM cancers were associated with higher levels of cancer pathway genes sets such as immune response (TNFA signaling via NFKB, interferon alpha, gamma response), apoptosis, hypoxia, reactive oxygen species, K-Ras and p53 signaling, whereas non-AAM tumors were associated with higher levels of fatty acid metabolism, glycolysis, Myc targets, mitotic spindle, DNA repair, PI3K via AKT/mTOR and WNT via beta-catenin signaling.
In this work, the authors conclude that significant biological differences in prostate cancer appear to be dependent in part upon a man’s racial ancestry. AAM patients had higher Decipher scores in low-grade tumors, higher expression of SPINK1, immune-related pathways, lower Decipher scores in higher grade tumors, lower expression of androgen receptor, androgen receptor downstream targets, ERG and DNA repair. Additional ancestry and race data should be collected from African American as well as other racial groups to further study its impact on the heterogeneity of prostate cancer and potential implications for treatment in the future.
Figure 1: Racial variation of molecular subtypes:
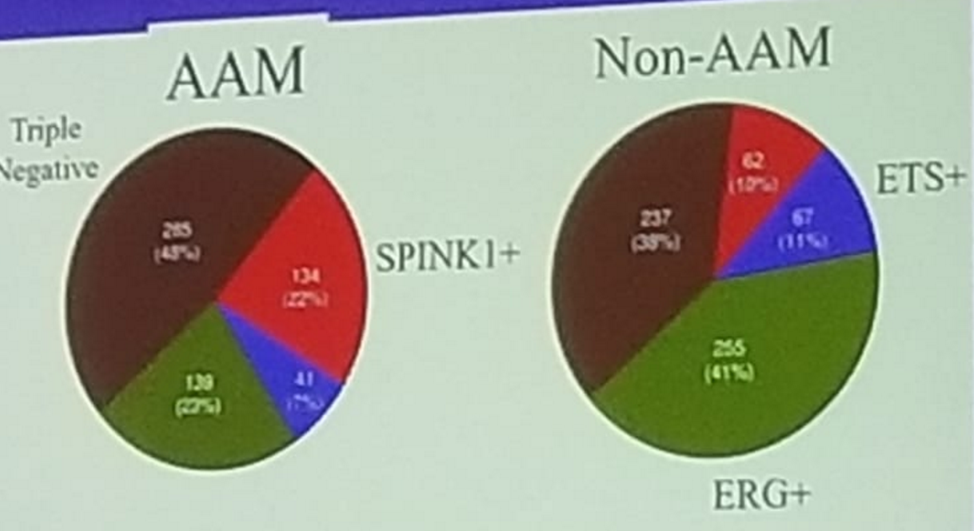
Presented by: Walter Rayford, MD, The Urology Group Memphis, TN, USA
Co-Authors: Jordan Alger, Washington, DC, Mandeep Takhar, Jennifer Jordan, Vancouver, Canada, Kamlesh Yadav, New York, NY, Mohammed Alshalalfa, Darlene Dai, Nick Fishbane, Takara Scott, Jill Collins, Britney Bender, Vancouver, Canada, Mohsen Ahmed, New York, NY, Jonathan Lehrer, Nicholas Erho, Elai Davicioni, Vancouver, Canada, Mark D. Greenberger, Memphis, TN, Randy Bradley, Knoxville, TN, Wei Phin Tan, Chicago, IL, Justin Watson, Marietta, GA, Mathew Beamer, Lambros Stamatakis, Washington, DC, Darrell J Carmen, Riverdale, GA, Kosj Yamoah, Tampa, FL, Jonathan Hwang, Washington, DC, Leslie A. Deane, Chicago, IL, Ashutosh Tewari, New York, NY
Written by: Hanan Goldberg, MD, Urologic Oncology Fellow (SUO), University of Toronto, Princess Margaret Cancer Centre @GoldbergHanan at the 2018 AUA Annual Meeting - May 18 - 21, 2018 – San Francisco, CA USA


