Urinary tract infections are the most common type of bacterial infection,1 accounting for at least 11 million physician office visits, 2 to 3 million emergency department visits, 400,000 hospitalizations, and approximately $2.3 billion in healthcare costs annually in the United States.2,3,4,5
This formidable burden on patients and the healthcare system continues to grow—for example, trends in population aging and the proliferation of antimicrobial-resistant bacteria are amplifying the need for intensive care and in-hospital mortality due to urinary tract infections.6,7
Such factors heighten the need to diagnose urinary tract infections and optimize antimicrobial therapy efficiently and accurately. However novel testing methods and our emerging understanding of the urinary microbiome raise questions about the relevance of traditional urine culture. In this article, I review major types of urinary tract infections, current approaches to the use of urine culture and sensitivity testing, recent evidence for the urinary microbiome, novel commercially available tests, and future approaches to the detection of urinary tract infections, including novel biomarkers and machine learning algorithms.
Uncomplicated versus complicated urinary tract infections
Distinguishing between uncomplicated and complicated urinary tract infections is an essential first step to guide management. Uncomplicated urinary tract infections are one of the most commonly treated types of infections in primary care settings. Women are at greatest risk, with a lifetime incidence approaching 50% and an approximately 33% rate of recurrence.8
The majority of patients with uncomplicated urinary tract infection are premenopausal women who are not pregnant and have few or no comorbidities. These patients typically present with symptoms of cystitis, such as dysuria, frequency, and urgency.9 Suprapubic pain and hematuria are uncommon, and pyelonephritis is not present.
Complicated urinary tract infections typically affect patients of either sex who have structural or functional abnormalities of the urinary tract.9 These patients often have had prior urologic procedures, recent antibiotic exposure, recent or long-term catheterization, or recent or current hospitalization (hospital-acquired urinary tract infections). Other high-risk groups include pregnant women, patients with diabetes mellitus, and patients with other immunocompromising conditions.
Patients with complicated urinary tract infections typically present with symptoms of pyelonephritis, including fever, chills, and flank pain, with or without nausea. They may have a confirmed or suspected history of infection(s) with more virulent bacteria. Urine cultures may reveal diverse microbiota and above-average patterns of antimicrobial resistance. Unfortunately, both these factors can undermine the efficacy of antimicrobial therapy.
When to consider urine culture and sensitivity
Historically, the diagnostic gold standard for urinary tract infections was to perform a standard urine culture (that is, spread 1 microliter of midstream urine onto 5% sheep blood and McConkey agars and incubate them aerobically).1 In this context, urinary tract infection was defined as the presence of an isolated, known uropathogen at a concentration of >105 CFU/ml or >102 CFU/ml in a symptomatic patient. Given the high prevalence of urinary tract infections and the fact that a urine culture is not a point-of-care test, it is worth considering when it is possible to forego culture without compromising treatment outcomes.
Current guidelines from the Infectious Diseases Society of America (IDSA) do not recommend routine standard urine culture for patients suspected to have uncomplicated urinary tract infections.10 There is considerable evidence that urinalysis is informative in this setting. In one study, approximately 94% of patients with a negative urinalysis also had a negative culture.11 In a longitudinal retrospective study of nearly 21,000 female outpatients, a negative urinalysis was associated with a 2.5-fold greater probability of a negative culture (<103 CFU/ml) compared with a positive urinalysis.12 For patients with recurrent symptoms of urinary tract infection, a prior negative urinalysis or urine culture or current vaginal irritation or discharge were associated with statistically significant increases in the probability of a negative culture.
Leukocyte esterase is especially informative when evaluating suspected uncomplicated urinary tract infections. In a large retrospective study of more than 8,500 such patients, a negative result for leukocyte esterase predicted a negative urine culture result with 95% accuracy (negative predictive value, 0.95), while negative results for both leukocyte esterase and nitrate were only slightly more accurate (combined negative predictive value, 0.96).13 Of note, a negative nitrate result alone was markedly less reliable for predicting a negative culture result (negative predictive value, 0.87).
For patients with uncomplicated urinary tract infections, the IDSA guidelines recommend selecting among the following drugs for empiric antimicrobial therapy:10
• Nitrofurantoin 100mg twice daily (BID) for 5 days
• Trimethoprim/sulfamethoxazole (Bactrim DS) for 3 days if local resistance rates are <20% (if you do not already have access to these data, consider contacting local, county, or state health departments)
• Fosfomycin (single 3-gram dose)
• Note that fluoroquinolones such as ciprofloxacin are no longer recommended for uncomplicated urinary tract infections due to high rates of resistance in some areas
In addition, the selection of antimicrobial therapy for uncomplicated urinary tract infections should guided by local patterns of antimicrobial resistance and the clinician’s best, informed guess as to the most likely uropathogen. In a study of more than 9,000 women with culture-confirmed uncomplicated urinary tract infections, 19% of isolates were resistant to trimethoprim-sulfamethoxazole and 12% were resistant to nitrofurantoin (10% were resistant to ciprofloxacin, which is no longer recommended for empiric therapy).14 Significant predictors of antimicrobial resistance included living in a ZIP code with above-average rates of antimicrobial resistance, having a history of infection with a resistant uropathogen, or having been prescribed antimicrobial therapy within the past 2 years. Based on these data, the investigators developed an algorithm that selected appropriate antimicrobial therapy in 92% of cases, exceeding the provider choice rate of 87.5%.
In conclusion, for patients with uncomplicated urinary tract infections, a traditional urine culture is not obsolete, but in most cases, it is unnecessary.
For patients suspected to have complicated urinary tract infections, clinicians should select initial (empiric) antimicrobial therapy on historical culture and sensitivity data, if available, in addition to the current results of urinalysis and local patterns of antimicrobial resistance. If the urinalysis is negative, and the patient has a prior history of negative cultures, then there is an 87% likelihood that the present symptoms are not caused by a urinary tract infection, according to recent data.12 These patients should be evaluated for other causes of their symptoms, such as chronic pelvic pain syndrome (CPPS) or tumors of the urinary tract. If the urinalysis is positive, then I recommend treating empirically and ordering a urine culture and sensitivity in order to modify treatment if necessary.
Urine is not sterile
Until very recently, the paradigm for managing both uncomplicated and complicated infections was to eliminate microorganisms from the urinary tract. Experts now recognize that this approach is limited by two key related factors: microbiota are present in the asymptomatic (healthy) urinary tract, and standard culture is relatively insensitive for the detection of urinary microorganisms.
Robust and mounting evidence for the existence of a diverse urinary microbiome challenges the old clinical maxim that “urine is sterile.”15 In one recent study, researchers performed quantitative PCR of 16S ribosomal RNA from 16 urine specimens from healthy men and women aged 26 to 90 years.16 Five samples tested positive for one bacterial phylum and one to six genera, while the rest contained an average of five phyla and eight to 36 genera. In another study, researchers performed high-throughput PCR sequencing of urine samples from eight women, all of whom had negative standard urine cultures.17 All samples were polymicrobial and showed considerable inter-specimen variability: 45 unique bacterial species were identified, of which nine were associated with urinary tract infections and 20 were of unknown pathogenic potential. Similar studies have confirmed this finding,18 suggesting that urine from healthy individuals frequently contains bacteria that standard urine cultures do not detect.
Another study delved deeper by using enhanced quantitative urine culture (EQUC) to determine whether bacteria identified in urine specimens were viable.19 This method involves plating larger volumes of urine, incubating specimens under a wider range of growth conditions, and using longer incubation periods. Among 65 urine specimens evaluated by EQUC, 80% grew out bacteria, of which 92% were not detected by standard culture. Fully 35 genera were identified, of which Lactobacillus, Corynebacterium, Streptococcus, Actinomyces, and Staphylococcus were most common. Most bacterial species replicated to numbers below the detection threshold of standard urine culture protocols. These findings were confirmed by another recent study of 150 adults (half reporting urinary symptoms) in which the use of EQUC detected 182 uropathogens—three times the number detected by standard culture.20
Collectively, the results of these studies confirm that diverse urinary microbiota are present both in healthy individuals and in patients with urinary symptoms, and that standard urine culture methods are relatively insensitive for characterizing this microbiome, including uropathogens. Standard cultures also do not reliably simulate biofilms, such as those formed in urinary catheters, and the final results of standard culture and sensitivity assays require days to be reported.11,21 Clearly, there is room for improvement.
Novel, commercially available tests for urinary tract infections
Three commercially available urine tests detect uropathogens with greater sensitivity than standard urine culture and also rapidly screen for evidence of antimicrobial resistance (Table 1).18,19,22 Potential benefits of these tests include faster pathogen identification, more accurate sensitivity results, and individualized treatment, all of which can theoretically improve patient outcomes.23,24,25
The Vikor Urine-ID™ test is a PCR-based assay that is able to detect multiple species of bacteria and also can identify up to 30 bacterial genes encoding resistance to antimicrobial agents.26 test results are available in approximately 12 to 24 hours (Table 1). Together with the test results, the manufacturer provides information on regional antimicrobial sensitivity patterns, the spectra of activity of antibiotics, medication costs, and relevant guidance from the U.S. Food and Drug Administration (FDA).
The MicroGenDX UTI Urinalysis Kit is a two-step test.27 First, quantitative PCR is used to screen for 16 types of bacteria as well as Candida albicans, with results provided within 24 hours. Second, next-generation sequencing (NGS) is used to compare microbiota DNA from the urine specimen with a database of more than 30,000 microbial species, with results in 3 to 5 days (Table 1). According to the manufacturer, the UTI Urinalysis Kit identifies all microbes in a urine specimen with 99.9% accuracy.
Table 1. Overview of novel commercially available tests for urinary tract infections
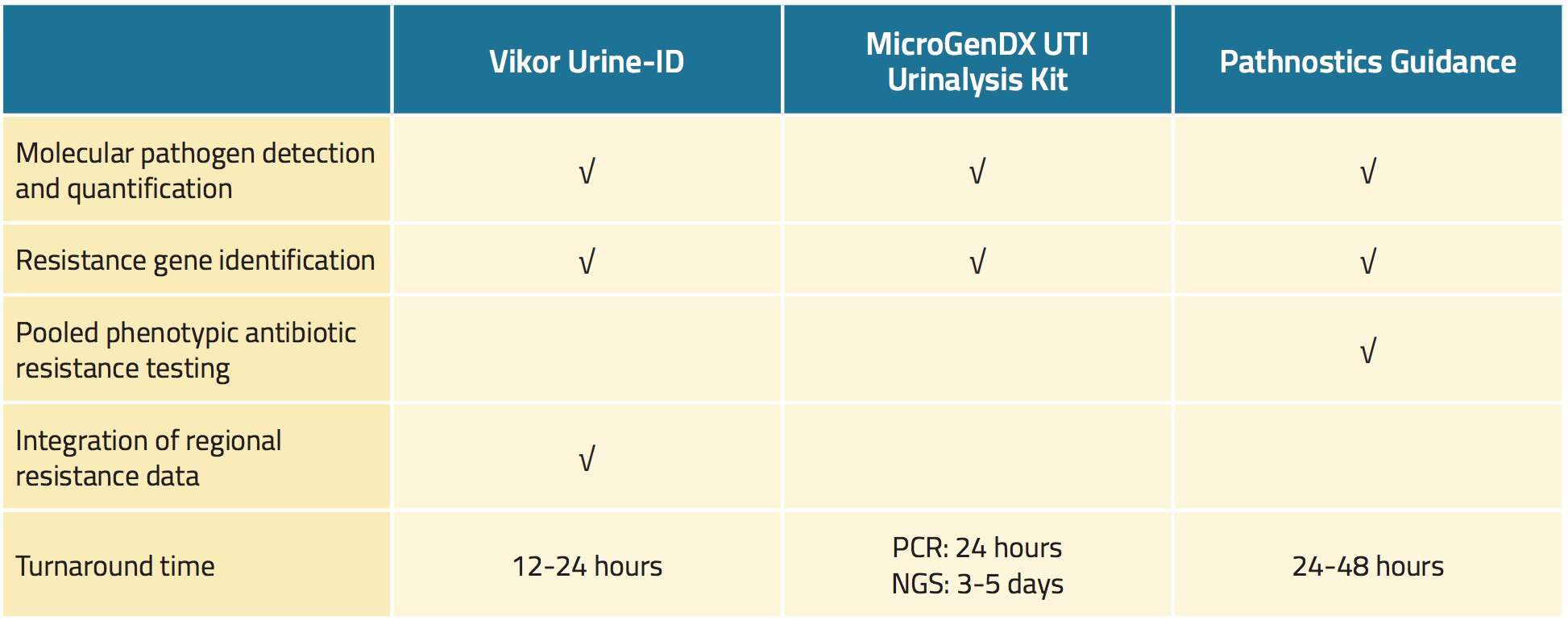
The Pathnostics Guidance assay consists of a suite of novel urine tests for patients with prostatitis, interstitial cystitis, and recurrent urinary tract infections.28 The assay for tract infection utilizes PCR to screen for 45 pathogens and 38 resistance genes. The Pathnostics Guidance assay also generates phenotypic data and pooled antimicrobial sensitivity results. The test can use either voided or catheter urine that can be stored at room temperature for up to 5 days. Results are available in 24 to 28 hours.
One caveat warrants mention. As we have discussed, these highly sensitive assays can detect bacteria in the urine of asymptomatic individuals and persons whose symptoms are unrelated to urinary tract infections. Therefore, clinicians should not base their treatment decisions on test results alone. Doing so can have adverse consequences for patients. In one study of 673 young and middle-aged women with asymptomatic bacteriuria and a history of recurrent urinary tract infections, half of patients received antimicrobial therapy while the rest did not.22 At 6-month and 12-month follow-up, antimicrobial therapy was associated with statistically significant increases in rates of urinary symptom recurrence. To avoid the overuse of antimicrobial agents and their associated side effects, costs, and selection for antimicrobial resistance, it is vital that clinicians evaluate test results in the context of patients’ overall risk and history of urinary tract infections and current clinical presentation.
Future directions for urine-based testing
Even as recently approved tests are poised to substantially improve the detection and management of urinary tract infections compared with standard culture alone, researchers continue to seek ways to improve diagnosis, pathogen detection, the quantification of antimicrobial sensitivity, and treatment algorithms. Examples include the use of machine learning techniques, composite analyses of novel biomarkers, and tests for cellular protein signatures.
Machine learning algorithms have shown early promise for improving the detection and evaluation of patients with uncomplicated urinary tract infections. For example, one study evaluated 17 clinical variables and 42 immunologic variables to identify the best predictors of the results of urine culture. Urine cloudiness (turbidity) was the best clinical predictor, with a positive predictive value of 0.65 (meaning that the presence of urine turbidity identified a culture-positive specimen with 65% accuracy) and a negative predictive value of 0.79 (meaning that the absence of urine turbidity identified a culture-negative specimen with 79% accuracy).29 Hence, urine turbidity was relatively insensitive but showed reasonable specificity for the detection of a culture-positive specimen. Together, four urinary biomarkers—matrix metallopeptidase 9, neutrophil gelatinase-associated lipocalin, interleukin (IL)-8, and IL-1β—achieved a substantially higher positive predictive value of 0.82, and a comparable negative predictive value of 0.76. Although combining these four urinary biomarkers with urine turbidity did not further improve these predictive values, the study results indicated that machine learning algorithms can reliably identify most patients with uncomplicated urinary tract infections.29
As machine learning models become more refined, their predictive values might improve, which might further improve the diagnosis and treatment of urinary tract infections in vulnerable populations while simultaneously reducing diagnostic workloads. For example, a recent large study analyzed more than 212,000 reports of urine microscopy, culture, and sensitivity results from three hospitals and outpatient services in Britain.30 Specific machine learning algorithms were designed for high-risk subgroups, such as pregnant women, children, and individuals with persistent or recurrent urinary tract infections. Machine learning detected approximately 95% of culture-positive specimens while reducing the workload associated with urine culture by approximately 41% and achieving approximately 24% higher specificity than a heuristic model based on white blood cell and bacterial counts. These are results are especially noteworthy in an era when population aging and the emergence of antimicrobial-resistant bacteria heighten the need to efficiently and accurately detect urinary tract infections that require treatment.
Recurrent urinary tract infections (historically defined as at least two urinary tract infections in the past 6 months, or more than three infections within the past year) have been found to affect approximately 50% among women over the age of 55 years and 27% of younger females.31, 32 Current guidelines call for intensive management with measures such as prophylactic antimicrobial therapy, limiting spermicide use, voiding after intercourse, immunoactive prophylaxis with OM-89, vaginal vaccination with Urovac, and the use of vaginal estrogen creams or rings in postpartum women.31, 33 Analyses of serum and urinary biomarkers might help better predict which patients will develop recurrent urinary tract infections so that they receive earlier, targeted interventions.32 Relevant biomarkers for recurrent urinary tract infections include decreased serum levels of vitamin D and prostate-specific antigen, increased serum levels of immunoglobulins, granulocyte colony-stimulating factor, macrophage colony-stimulating factor, and IL-5, while relevant urinary biomarkers include elevated levels of IL-8 and decreased levels of nerve growth factor and neutrophil gelatinase-associated lipocalin.
Future diagnostics might also include point-of-care biomarker assays that supplement or reduce the need for urine cultures. An ideal assay would be as rapid and inexpensive as a urine dipstick test, but more precise and accurate. Potential candidates include trimethylamine and acetate, which are markers of bacterial metabolism, and xanthine oxidase and myeloperoxidase, which are enzymatic biomarkers.34
Future assays might also evaluate the urinary exosome as a potential biomarker for urinary tract infections. The exosome consists of small extracellular vesicles that carry cellular proteins. In one study, the urinary proteins Akt (an intracellular signaling protein) and CD9 (an intracellular transmembrane protein) were significantly elevated in women with urinary tract infections compared with women with asymptomatic bacteriuria, and levels of both proteins significantly decreased following antimicrobial therapy.35 The results of such studies could someday facilitate the development of commercial assays that evaluate the urinary exosome to help guide treatment decision-making.
Summary
The evaluation and management of urinary tract infections is integral to urology practice. For uncomplicated urinary tract infections, urine culture is typically unnecessary, and empiric treatment according to current IDSA guidelines is appropriate. Empiric therapy based on historical culture and sensitivity results is also appropriate for complicated urinary tract infections but should be modified based on current culture and sensitivity. Recurrent urinary tract infections may warrant require intensive therapeutic and behavioral interventions.
While these are our current best practices, they leave substantial room for improvement, particularly considering recent evidence for the existence of a urinary microbiome and the limitations of standard screening and culture. There is a need for rapid, reliable assays to better detect uropathogens, differentiate clinically meaningful urinary dysbiosis from clinically inconsequential bacteriuria, and guide antimicrobial therapy based on personalized resistance data. Sensitivity assays based on quantitative PCR and next-generation sequencing are now available. In the future, the use of enhanced quantitative urine culture methods may also improve clinical care. Investigators also are evaluating machine learning algorithms that incorporate serum and urinary biomarkers and other clinical variables to more effectively detect urinary tract infections, predict recurrence, and reduce the diagnostic workload. Incorporating these novel approaches with local resistance and hospital readmission data may further their utility. The end goal is to develop sensitive, specific, cost-effective tests and practical algorithms that improve management and patient outcomes.
Written by: Edward Schaeffer, MD, Ph.D., Chair, Department of Urology, Feinberg School of Medicine, Program Director, Genitourinary Oncology Program, Robert H. Lurie Comprehensive Cancer Center, Northwestern University, Chicago, Illinois
References:
1. Schaeffer, AJ., Matulewicz, RS., & Klumpp, DJ. (2016). Infections of the Urinary Tract. In AJ. Wein, & et al (Eds.), Campbell-Walsh Urology, Eleventh Edition Philadelphia: Elsevier-Saunders.
2. Schappert, S. M., and E. A. Rechtsteiner. "Ambulatory medical care utilization estimates for 2007." Vital and Health Statistics. Series 13, Data from the National Health Survey 169 (2011): 1-38.
3. Foxman, Betsy. "Urinary tract infection syndromes: occurrence, recurrence, bacteriology, risk factors, and disease burden." Infectious disease clinics of North America 28, no. 1 (2013): 1-13.
4. Foxman, Betsy. "The epidemiology of urinary tract infection." Nature Reviews Urology 7, no. 12 (2010): 653.
5. Simmering, Jacob E., Fan Tang, Joseph E. Cavanaugh, Linnea A. Polgreen, and Philip M. Polgreen. "The increase in hospitalizations for urinary tract infections and the associated costs in the United States, 1998–2011." In Open forum infectious diseases, vol. 4, no. 1. Oxford University Press, 2017.
6. Nguyen, Hoa Q., Nga TQ Nguyen, Carmel M. Hughes, and Ciaran O’Neill. "Trends and impact of antimicrobial resistance on older inpatients with urinary tract infections (UTIs): A national retrospective observational study." PloS one 14, no. 10 (2019).
7. Critchley, Ian A., Nicole Cotroneo, Michael J. Pucci, and Rodrigo Mendes. "The burden of antimicrobial resistance among urinary tract isolates of Escherichia coli in the United States in 2017." PloS one 14, no. 12 (2019).
8.Gupta, Kalpana, and Barbara W. Trautner. "Diagnosis and management of recurrent urinary tract infections in non-pregnant women." Bmj 346 (2013): f3140.
9. Tan, Chee Wei, and Maciej Piotr Chlebicki. "Urinary tract infections in adults." Singapore medical journal 57, no. 9 (2016): 485.
10. Gupta, Kalpana, Thomas M. Hooton, Kurt G. Naber, Björn Wullt, Richard Colgan, Loren G. Miller, Gregory J. Moran et al. "International clinical practice guidelines for the treatment of acute uncomplicated cystitis and pyelonephritis in women: a 2010 update by the Infectious Diseases Society of America and the European Society for Microbiology and Infectious Diseases." Clinical infectious diseases 52, no. 5 (2011): e103-e120.
11. Huang, Bin, Lei Zhang, Weizheng Zhang, Kang Liao, Shihong Zhang, Zhiquan Zhang, Xingyan Ma et al. "Direct detection and identification of bacterial pathogens from urine with optimized specimen processing and enhanced testing algorithm." Journal of clinical microbiology 55, no. 5 (2017): 1488-1495.
12. Cohen, Jason E., Emily M. Yura, Liqi Chen, and Anthony J. Schaeffer. "Predictive Utility of Prior Negative Urine Cultures in Women with Suspected Recurrent Uncomplicated Urinary Tract Infections." The Journal of urology 202, no. 5 (2019): 979-985.
13. Marques, Alexandre Gimenes, Jacyr Pasternak, Márcio dos Santos Damascena, Carolina Nunes França, and Marinês Dalla Valle Martino. "Performance of the dipstick screening test as a predictor of negative urine culture." Einstein (São Paulo) 15, no. 1 (2017): 34-39.
14. Cohen, Jason E., Liqi Chen, and Anthony J. Schaeffer. "Algorithms Using Previous Resistance, Prior Antimicrobial Prescriptions, and Patient Place of Residence Enhance Empirical Therapy for Women With Uncomplicated Urinary Tract Infections." Urology 137 (2020): 72-78.
15. Finucane, Thomas E. "‘Urinary tract infection’ and the microbiome." The American journal of medicine 130, no. 3 (2017): e97-e98.
16. Lewis, Debbie Ann, Richard Brown, Jon Williams, Paul White, Susan Kim Jacobson, Julian Marchesi, and Marcus John Drake. "The human urinary microbiome; bacterial DNA in voided urine of asymptomatic adults." Frontiers in cellular and infection microbiology 3 (2013): 41.
17. Siddiqui, Huma, Alexander J. Nederbragt, Karin Lagesen, Stig L. Jeansson, and Kjetill S. Jakobsen. "Assessing diversity of the female urine microbiota by high throughput sequencing of 16S rDNA amplicons." BMC microbiology 11, no. 1 (2011): 244.
18. Wolfe, Alan J., Evelyn Toh, Noriko Shibata, Ruichen Rong, Kimberly Kenton, MaryPat FitzGerald, Elizabeth R. Mueller et al. "Evidence of uncultivated bacteria in the adult female bladder." Journal of clinical microbiology 50, no. 4 (2012): 1376-1383.
19. Hilt, Evann E., Kathleen McKinley, Meghan M. Pearce, Amy B. Rosenfeld, Michael J. Zilliox, Elizabeth R. Mueller, Linda Brubaker, Xiaowu Gai, Alan J. Wolfe, and Paul C. Schreckenberger. "Urine is not sterile: use of enhanced urine culture techniques to detect resident bacterial flora in the adult female bladder." Journal of clinical microbiology 52, no. 3 (2014): 871-876.
20. Price, Travis K., Tanaka Dune, Evann E. Hilt, Krystal J. Thomas-White, Stephanie Kliethermes, Cynthia Brincat, Linda Brubaker, Alan J. Wolfe, Elizabeth R. Mueller, and Paul C. Schreckenberger. "The clinical urine culture: enhanced techniques improve detection of clinically relevant microorganisms." Journal of clinical microbiology 54, no. 5 (2016): 1216-1222.
21. Sathiananthamoorthy, Sanchutha, James Malone-Lee, Kiren Gill, Anna Tymon, Trang K. Nguyen, Shradha Gurung, Linda Collins et al. "Reassessment of routine midstream culture in diagnosis of urinary tract infection." Journal of clinical microbiology 57, no. 3 (2019): e01452-18.
22. Cai, Tommaso, Sandra Mazzoli, Nicola Mondaini, Francesca Meacci, Gabriella Nesi, Carolina D'Elia, Gianni Malossini, Vieri Boddi, and Riccardo Bartoletti. "The role of asymptomatic bacteriuria in young women with recurrent urinary tract infections: to treat or not to treat?." Clinical infectious diseases 55, no. 6 (2012): 771-777.
23. Lehmann, Lutz E., Stefan Hauser, Thomas Malinka, Sven Klaschik, Stefan U. Weber, Jens-Christian Schewe, Frank Stüber, and Malte Book. "Rapid qualitative urinary tract infection pathogen identification by SeptiFast® real-time PCR." PLoS One 6, no. 2 (2011).
24. Schmidt, K., K. K. Stanley, R. Hale, L. Smith, J. Wain, J. O'grady, and D. M. Livermore. "Evaluation of multiplex tandem PCR (MT-PCR) assays for the detection of bacterial resistance genes among Enterobacteriaceae in clinical urines." Journal of Antimicrobial Chemotherapy 74, no. 2 (2019): 349-356.
25. Barczak, Amy K., James E. Gomez, Benjamin B. Kaufmann, Ella R. Hinson, Lisa Cosimi, Mark L. Borowsky, Andrew B. Onderdonk et al. "RNA signatures allow rapid identification of pathogens and antibiotic susceptibilities." Proceedings of the national academy of sciences 109, no. 16 (2012): 6217-6222.
26. Urine-IDTM – Vikor Scientific. https://www.vikorscientific.com/test-menu/urine-id/. Accessed February 15, 2020.
27. UTI Urine Kit - Insurance | MicroGen Diagnostics. https://microgendx.com/product/urine-kit-ins/. Accessed February 15, 2020.
28. Guidance - Pathnostics. https://www.pathnostics.com/physicians/tests/guidance/. Accessed February 15, 2020.
29. Gadalla, Amal AH, Ida M. Friberg, Ann Kift-Morgan, Jingjing Zhang, Matthias Eberl, Nicholas Topley, Ian Weeks et al. "Identification of clinical and urine biomarkers for uncomplicated urinary tract infection using machine learning algorithms." Scientific Reports 9, no. 1 (2019): 1-11.
30. Burton, Ross J., Mahableshwar Albur, Matthias Eberl, and Simone M. Cuff. "Using artificial intelligence to reduce diagnostic workload without compromising detection of urinary tract infections." BMC medical informatics and decision making 19, no. 1 (2019): 171.
31. Bonkat, G., R. Pickard, R. Bartoletti, F. Bruyere, S. E. Geerlings, and F. Wagenlehner. "Guidelines on urological infections. EAU Guidelines." European Association of Urology (2017).
32. Jhang, Jia-Fong, and Hann-Chorng Kuo. "Recent advances in recurrent urinary tract infection from pathogenesis and biomarkers to prevention." Tzu-Chi Medical Journal 29, no. 3 (2017): 131.
33. Dason, Shawn, Jeyapandy T. Dason, and Anil Kapoor. "Guidelines for the diagnosis and management of recurrent urinary tract infection in women." Canadian Urological Association Journal 5, no. 5 (2011): 316.
34. Karlsen, H., and T. Dong. "Biomarkers of urinary tract infections: state of the art, and promising applications for rapid strip-based chemical sensors." Analytical Methods 7, no. 19 (2015): 7961-7975.
35. Mizutani, Kosuke, Kyojiro Kawakami, Kengo Horie, Yasunori Fujita, Koji Kameyama, Taku Kato, Keita Nakane et al. "Urinary exosome as a potential biomarker for urinary tract infection." Cellular microbiology 21, no. 7 (2019): e13020.
Related Content:
Download: Everyday Urology: Volume 5, Issue 1


