PROfound is a global study with patients screened from 20 countries including the USA, Europe, and Asia. To screen patients for PROfound, an investigational next-generation sequencing assay was used to prospectively select patients harboring homologous recombination repair gene mutations in their tumor tissue. This tumor tissue test reports deleterious or suspected deleterious alterations in 15 homologous recombination repair genes. Tissue samples were clinically heterogeneous regarding location and timing. To be eligible for PROfound, patients had to have a qualifying mutation and meet the remaining eligibility requirements prior to being randomized. The study comprised two cohorts:
- Cohort A - patients with mutations in BRCA1, BRCA2 or ATM (assigned regardless of any co-occurring mutation in other genes)
- Cohort B - 12 other homologous recombination repair genes
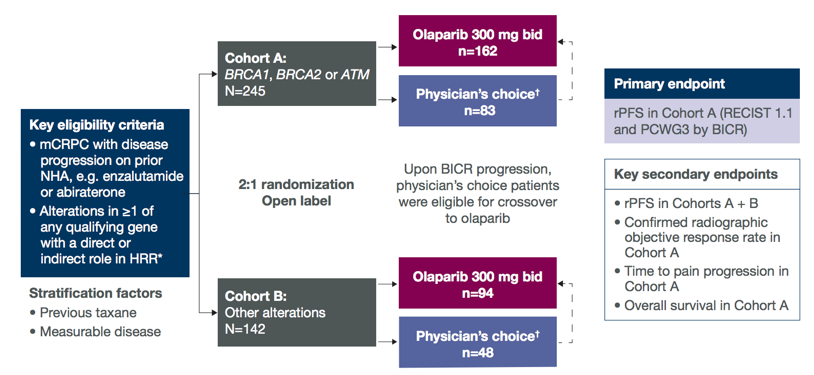
Among the 4,425 screened patients, 4,047 had samples tested of whom 2,792 (69%) yielded an interpretable result. Most archived samples (79.7%) were from the primary tumor and 10.1% were derived from metastatic tissue. Samples were derived from newly collected tissue in 9.0% of patients, 3.4% from the primary and 5.6% from the metastatic biopsy. A qualifying homologous recombination repair gene mutation was detected in 778 (27.9%) patients and the prevalence of homologous recombination repair gene mutations in genes included in Cohort A was 17.1%. The most common gene mutation was BRCA2 (8.7%).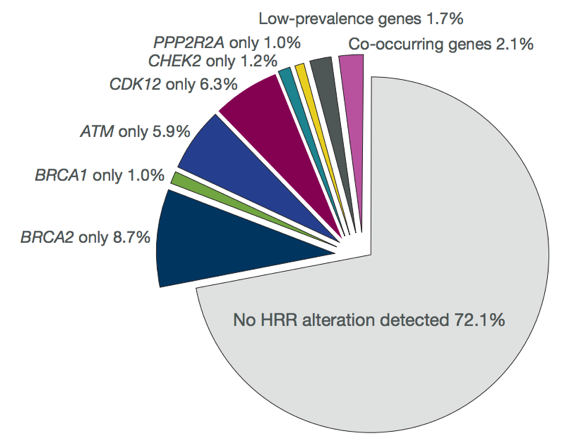
A co-occurring qualifying homologous recombination repair gene alteration in ≥ 1 gene was detected in 59 (7.6%) patients, most commonly BRCA2 (n = 30), CDK12 (n = 24) or ATM (n = 13).
Dr. De Bono’s concluding statements assessing the gene mutation profile among men screened for PROfound is as follows:
- This is the largest study to date with central prospective homologous recombination repair gene mutation tissue testing in prostate cancer
- A qualifying homologous recombination repair gene mutation was observed in 27.9% of mCRPC patients with a successful tumor test result
- Homologous recombination repair gene mutations were most common in BRCA2, followed by ATM and CDK12
- Further analyses are warranted to help understand if there is any correlation between clinical/demographic characteristics and homologous recombination repair gene mutations.
- This study confirms that a significant proportion of mCRPC patients with alterations in homologous recombination repair genes can be identified through tissue testing
Clinical Trial Identification: NCT02987543
Presented by: Johann S. De Bono, MB ChB, FRCP, MSc, PhD, Head of the Division of Clinical Studies, Professor in Experimental Medicine, The Institute of Cancer Research (ICR), Royal Marsden Hospital, London, UK
Co-Authors: J. De Bono1, K. Fizazi2, F. Saad3, N. Shore4, S. Sandhu5, N. Mehra6, M. Kolinsky7, G. Roubaud8, M. Ӧzgüroǧlu9, N. Matsubara10, C. Gedye11, Y.D. Choi12, C. Padua13, C. Goessl14, A. Kohlmann15, C. Corcoran16, C. Adelman17, A. Allen18, J. Burgents19, M. Hussain20
1. Institute of Cancer Research, London, UK
2. Institut Gustave Roussy, Villejuif, FR
3. Hospital St. Luc du CHUM, Montreal, CA
4. Medical Director, Myrtle Beach, US
5. Peter MacCallum Cancer Centre, Melbourne, A
6. Radboud University Medical Center, Nijmegen, NL
7. University of Alberta Cross Cancer Institute, Edmonton, CA
8. Institute Bergonié, Bordeaux, FR
9. Istanbul University-Cerrahpaşa, Cerrahpaşa School of Medicine, Istanbul, TR
10. National Cancer Center East, Chiba, JP
11. School of Medicine and Public Health, University of Newcastle, Callaghan, AU
12. Yonsei University College of Medicine, Seoul, KR
13. Cetus Medicina Oncológica, Betim, BR
14. BN ImmunoTherapeutics, Inc., Mountain View, US
15. AstraZeneca UK, Royston, UK
16. AstraZeneca UK, Cambridge, UK
17. AstraZeneca, Cambridge, UK
18. AstraZeneca, Gaithersburg, US
19. Merck, Sharpe & Dohme NA, Kenilworth, US
20. Northwestern University Feinberg School of Medicine, Chicago, US
Written by: Zachary Klaassen, MD, MSc – Assistant Professor of Urology, Georgia Cancer Center, Augusta University/Medical College of Georgia, Twitter: @zklaassen_md at the 2019 European Society for Medical Oncology annual meeting, ESMO 2019 #ESMO19, 27 Sept - 1 Oct, 2019 in Barcelona, Spain
Reference:


