He began highlighting data from the PROfound and TRITON2 studies which examined the role of PARP inhibitors in patients with metastatic castration-resistant prostate cancer (mCRPC). As has been previously reported, the PROfound trial demonstrated significant improvements in rPFS and OS among men in cohort A with mutations in BRCA1, BRCA2, and ATM. Similarly, TRITON2 found benefit to rucacparib, though this was not a randomized comparison. Dr. Nelson highlighted that many other mutations were included in these studies though benefit of PARP inhibition was not demonstrated in these. Overall, he emphasized that while these trials provided evidence of substantial clinical benefit in an advanced patient population using a biomarker-driven study design and have changed clinical practice, there are significant limitations as these patients are not cured and many responses are not durable. Further, a large proportion of biomarker-positive patients failed to demonstrate benefit and the resulting approval was not congruent with the precision focus.
Dr. Nelson then discussed mechanisms to inform biomarkers, focusing on the idea of synthetic lethality with an emphasis on DNA damage repair. In patients with “error-prone” DNA repair as a result of DDR mutations, drug-induced PARP inhibition leads to a lack of DNA repair as a resultant cellular death.

DNA damage repair is a complex, coordinated process involving many related proteins which serve to signal DNA damage and coordinate repair. Despite the many related components, Dr. Nelson emphasized that the importance of these is not equal. Given these differences, it is important to consider the test characteristics of a biomarker.
Dr. Nelson then emphasized the importance of considering both the precision and accuracy of biomarkers. This is particularly relevant given the list of biomarkers utilized for each of olaparib (comprising 15 mutations) and rucaparib (only 2: BRCA1 and BRCA2). Given the shared mechanism of the two agents, such discrepancies do not make sense according to Dr. Nelson.
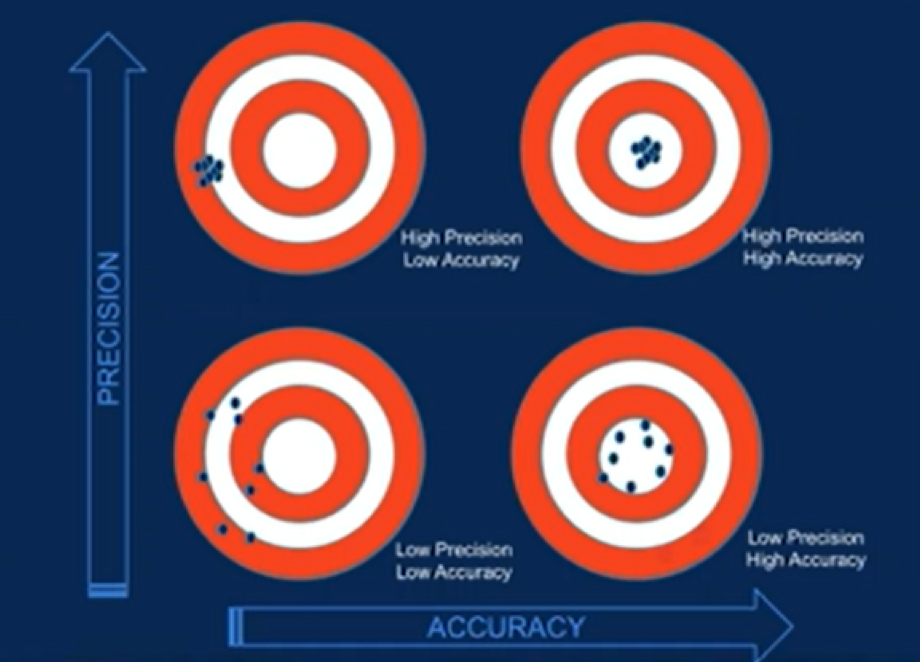
Further, differences in trial design and biomarker testing are unlikely to adequately explain differences in labeling and approval. Thus, in his view, the approved panel for olaparib lacks specificity while that for rucaparib lacks sensitivity. “Getting it wrong” with the appropriate use of biomarker-driven therapy risking administered ineffective therapy with associated toxicity and cost. Overall, he highlighted that approved biomarkers are limited by their indirect capture of homologous recombination repair deficiency, lack of mechanistic equivalence, and potential interference in assay interpretation. Further, there is a lack of evidence supporting many markers including ATM, CDK12, and CHEK2.
Examining the olaparib label specifically, the approved biomarkers make no mention of biallelic loss, include several markers which were never tested, and include several markers in which PARP inhibition has minimal response. The importance of biallelic loss is emphasized as this is the most predominant status in patients with ovarian cancer in which PARP inhibition has proven useful. However, this is less common in patients with prostate cancer, was not assessed (or at least not reported) in PROfound, and comprised only 36% of patients in TRITION2. Thus, current use would be associated with false-positive test results.
Dr. Nelson emphasized that the issue with gene mutational testing is that this indirectly reflect DNA repair deficiency. However, more important is functional deficiency. Assessment of functional deficiency may be identifiable based on genomic “scars.”
Using an example of a biomarker assay, Dr. Nelson emphasized that current testing has significant limitations which may limit their utility. CHIP variants (clonal hematopoiesis of indeterminant potential) may lead to false-positive identification of mutations in ctDNA which are neither germline nor somatically present in the tumor.
Moving forward, he emphasized the importance of refining assays and biomarker approvals as well as assessing PARP inhibition earlier in the disease course.
Presented by: Peter Nelson, MD, Fred Hutchinson Cancer Research Center
Written by: Christopher J.D. Wallis, Urologic Oncology Fellow, Vanderbilt University Medical Center Contact: @WallisCJD on Twitter during the 2021 American Society of Clinical Oncology Genitourinary Cancers Symposium (#GU21), February 11th-February 13th, 2021


