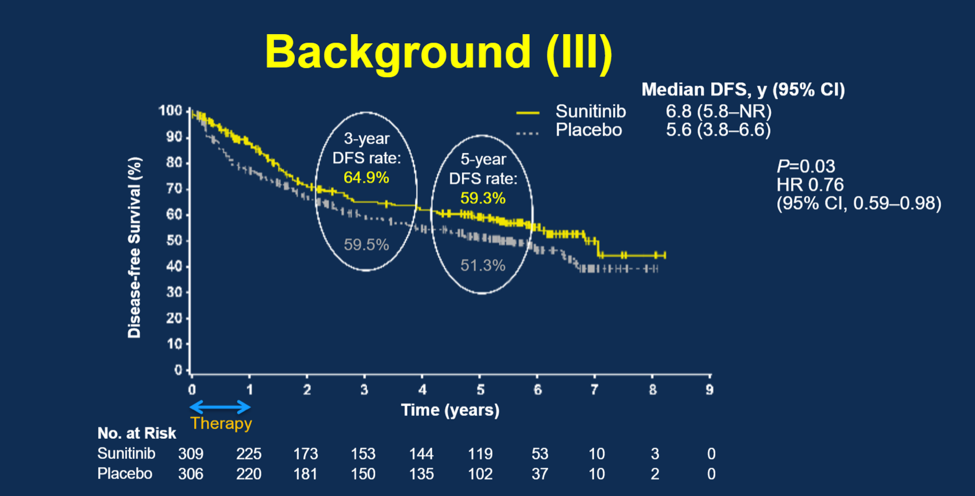
Dr. George and colleagues assessed the use of phamacogenomic biomarkers that might help to predict which patients responded to sunitinib. They evaluated 10 single nucleotide polymorphisms (SNPs) from patients in the S-TRAC trial and associated those SNPs with clinical outcomes (DFS and overall survival [OS]). They isolated 10 important SNPs that clustered around VEGF pathways or other key genes involved with angiogenesis and hypoxia.
3 SNPs (in VEGFR1, VEGFR2, and eNOS genes) significantly associated with DFS (HRs 0.44, 0.46, 0.53, respectively per SNP). These HRs are quite remarkable; demonstrating that these specific genotypes may help predict which patients will benefit the most from sunitinib adjuvant therapy. Importantly, they also identified a genotype that did demonstrably worse with sunitinib treatment, which could be an important biomarker for patients who may not respond well to sunitinib and should be stratified separately in future studies.
In conclusion, correlations between common SNPs in VEGFR1, VEGFR2, and eNOS genes and DFS in the S-TRAC population show that genomic biomarkers may be an important way to predict responses to adjuvant sunitinib. SNPs are easy to identify and type, so may be clinically meaningful markers. More importantly, these biomarkers may help stratify patients in future prospective studies to understand the true effect of these therapies in TKI-sensitive and TKI-resistant populations.
Presented by: Daniel J. George, MD, Duke Cancer Institute, Durham NC
Written by: Shreyas Joshi, MD, Fox Chase Cancer Center, Philadelphia, PA at the 2018 American Society of Clinical Oncology Genitourinary (ASCO GU) Cancers Symposium, February 8-10, 2018 - San Francisco, CA


