(UroToday.com) The 2022 American Society for Radiation Oncology (ASTRO) Annual Meeting held in San Antonio, TX between October 23rd and 26th, 2022 was host to a State of the Art session for the selection and treatment of localized prostate cancer. Karthik Meiyappan presented transcriptomic analysis of ATM expression in localized prostate cancer and its implications for radiotherapy treatment.
Ataxia Telangiectasia Mutated (ATM) gene is a DNA Damage and Repair (DDR) gene that is activated by radiation exposure via direct DNA damage and oxidative stress mechanisms. Mutations of this gene have been implicated in many cancers including Ataxia Telangiectasia, breast cancer, bladder cancer, and melanoma. ATM deficient or defective cells have been shown to have increased radiosensitivity. Prostate cancer patients with somatic or germline ATM mutations have been shown to have higher grade disease, lower PSA at diagnosis, and such mutations are present in 30 – 40% of mCRPC patients.1-4

Although there has been considerable work evaluating ATM mutations, to date there has been limited research evaluating ATM expression. As such, the objective of this study was to evaluate the possible relationship between ATM expression and other key DDR genes in a prospective cohort, with respect to:
- Metastasis-free survival
- Prostate cancer-specific mortality
- Determine whether innate ATM expression predicts a differential response to radiotherapy.
The authors hypothesized that higher ATM expression is a predictor of differential outcomes after radiotherapy.
To this end, the authors included patients from 3 cohorts:
- Decipher/GRID (n=5,239): A prospective cohort which included genome-wide expression profiles, had Decipher scores available, and had a median follow up of 48 months
- Mayo clinic (n=780): A retrospective cohort with a median follow up of 156 months. This cohort was used to assess MFS and PCSM.
- PORTOS/matched cohort (n=196): This cohort was stratified by radiotherapy versus no radiotherapy and was matched on pre-operative PSA, Gleason Score, margins, extraprostatic extension, seminal vesicle invasion, and lymph node involvement.
The baseline patient characteristics and previous treatments received by patients in these cohorts are demonstrated in the table below:
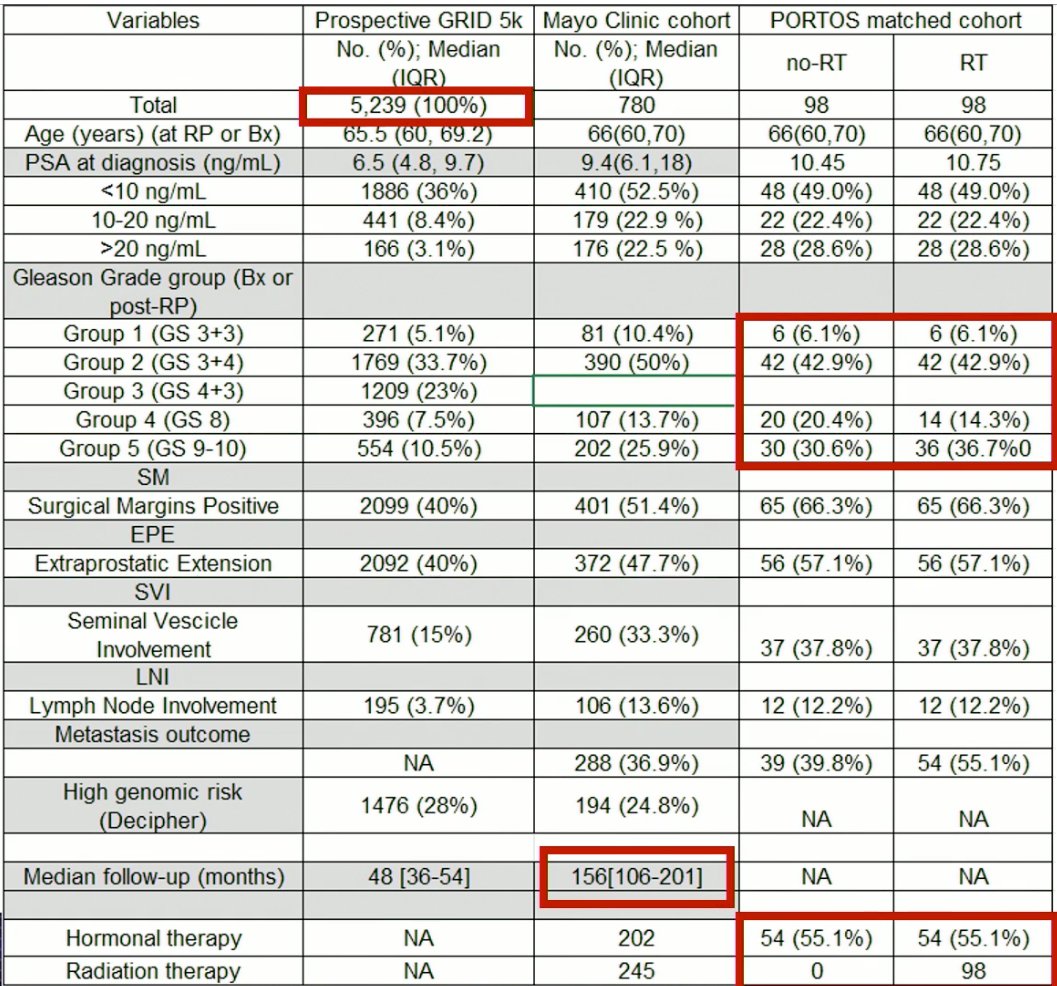
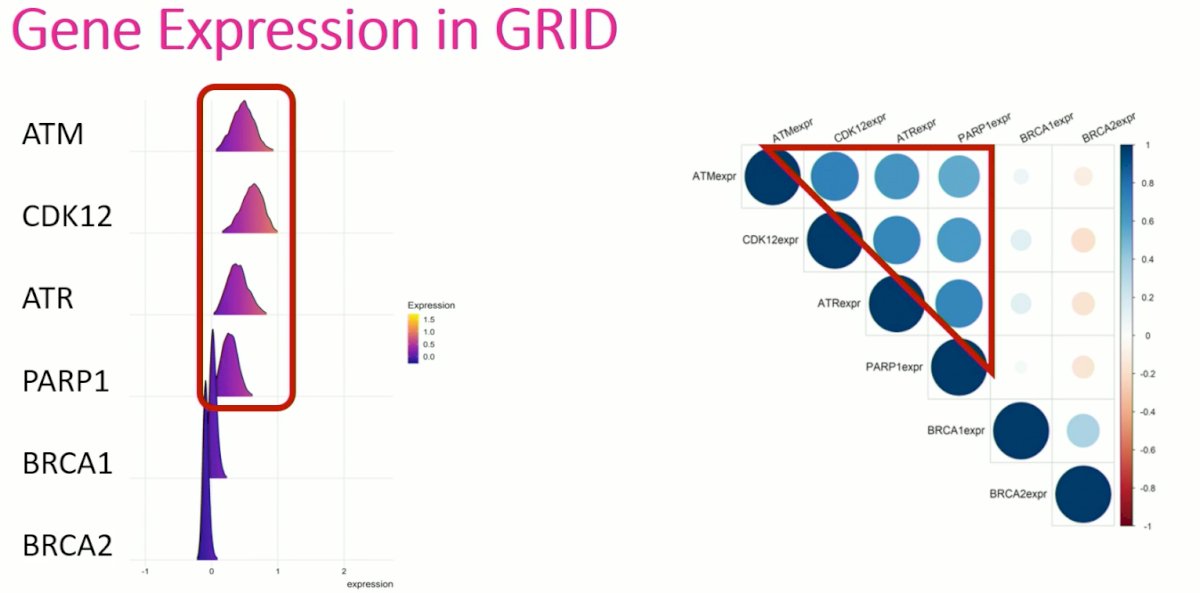
The authors first used the GRID cohort to evaluate gene expression of ATM, CDK12, ATR, PARP1, BRCA1, and BRCA2. The authors noted that gene expression was highly expressed for ATM, DK12, ATR, and PARP1, but not BRCA1/2. Furthermore, correlative studies demonstrated high co-expression between these 4 genes, but not with BRCA1/2.
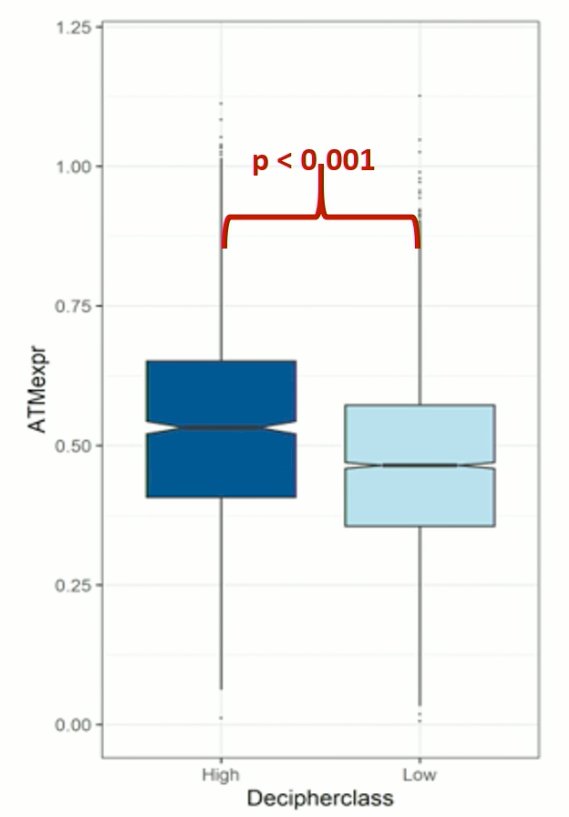
ATM was noted to be significantly higher expressed in the high Decipher score group, particularly significant given the known increased hazard of metastasis and prostate cancer-specific mortality in the high Decipher group patients.
Next, ATM expression was correlated with greater than 200 genomic signatures of outcome, biologic pathways and molecular subtypes was evaluated. ATM expression was noted to be highly correlated with numerous markers of angiogenesis, immune cell infiltration, and genomic instability.

Using data from the Mayo cohort, it was demonstrated that high ATM expression was associated with worse MFS and PCSM:

Does radiation treatment differentially affect outcomes in patients with high versus low ATM expression? Using data from the PORTOS matched cohort (all having received concurrent hormone therapy with adjuvant/salvage radiation), the authors found no significant differences in MFS in patients not receiving radiotherapy. However, in patients receiving adjuvant/salvage radiotherapy post-prostatectomy, there were significant differences by ATM expression levels, with patients with the highest quartile of ATM expression having significantly worse MFS.
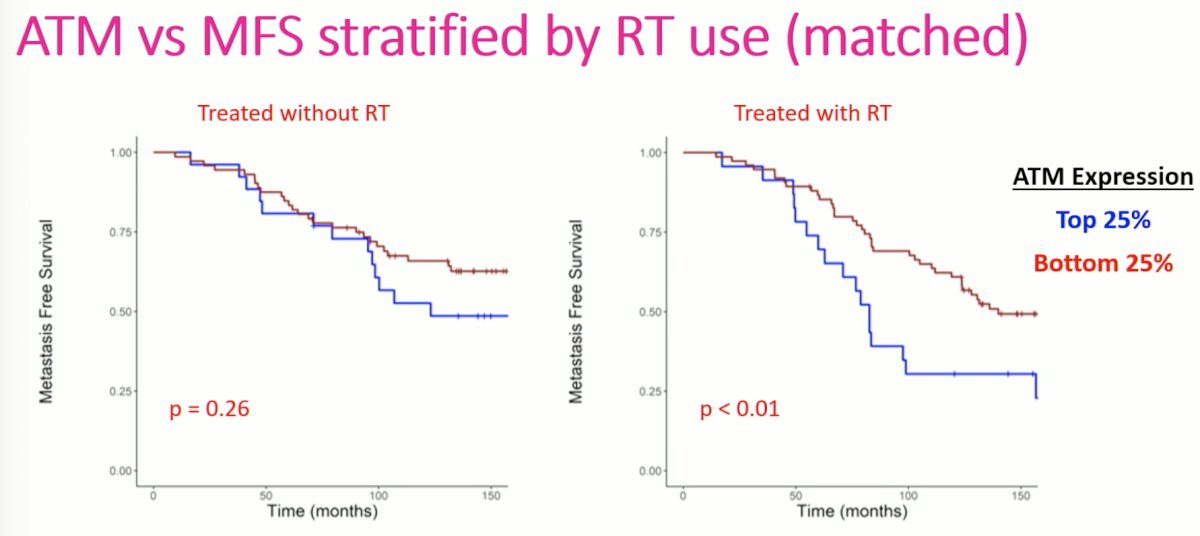
The authors discussed that these results suggest that ATM expression can act as a prognostic biomarker for MFS and PCSM, confirming preliminary findings observed in oligometastatic patients from the STOMP1 and ORIOLE2 trials. ATM expression can also act as a predictive biomarker for radiotherapy response. The authors postulated that the worse outcomes in patients with high ATM expression treated with radiation therapy may be due to improved DNA damage response, conferring radio-resistance, and thus increased hazard of disease progression and metastasis.
The presenter concluded his presentation with the following take home messages:
- ATM expression is highly correlated with several prostate cancer risk genes (CDK12, ATR, PARP1, FANCL) and genetic signatures of cancer progression (angiogenesis, immune infiltration, genomic instability).
- Higher ATM expression is a prognostic biomarker for worse MFS and PCSM.
- High ATM expression may be predictive of poor radiotherapy response in post-RP patients
Presented by: Karthik Meiyappan, BS, Medical Student, University of Miami Miller School of Medicine, Miami, FL @Dr_Karti_B
Written by: Rashid Sayyid, MD, MSc – Society of Urologic Oncology (SUO) Clinical Fellow at The University of Toronto, @rksayyid on Twitter during the 2022 American Society of Radiation Oncology (ASTRO) Annual Hybrid Meeting, San Antonio, TX, Sat, Oct 22 – Wed, Oct 26, 2022.
References:
- Ost P, Reynders D, Decaestecker K, et al. Surveillance or metastasis-directed therapy for oligometastatic prostate cancer recurrence: A prospective, randomized, multicenter phase II trial. J Clin Oncol. 2018;36(5):446-453.
- Phillips R, Shi WY, Deek M, et al. Outcomes of Observation vs Stereotactic Ablative Radiation for Oligometastatic Prostate CancerThe ORIOLE Phase 2 Randomized Clinical Trial. JAMA Oncol. 2020;6(5):650-659.

