(UroToday.com) On the second day of the American Society for Clinical Oncology (ASCO) Genitourinary Cancer Symposium 2022 focused on urothelial carcinoma, in Poster Session B, Dr. Grivas presented a poster examining the effect of PD-L1 expression, as determined by immunohistochemistry (IHC), on genomic alteration profiles in patients with urothelial bladder carcinoma (UBC).
To do so, they identified 102 cases of advanced UBC with known PD-L1 expression that subsequently underwent hybrid-capture-based comprehensive genomic profiling to evaluate all classes of genomic alterations. Tumor mutational burden (TMB) was determined on up to 1.1 Mbp of sequenced DNA and microsatellite instability (MSI) was determined on 114 loci. Tumor cell (TC) PD-L1 expression was determined by IHC (Dako 22C3).

Only patients with PD-L1 high (H) (≥50% TC expression) and negative (N) (0% TC expression) were included. Patients with PD-L1 Low (1-49% TC expression) were excluded from this study.
Among the 102 patients initially identified, 24 had high PD-L1 expression and 78 had negative PD-L1 expression.
Comparing patients with high and negative PD-L1 expression, only CD274 and FGFR3 rates varied significantly. Two (8.3%) of the 24 high PD-L1 expressing UBC featured CD274 (PD-L1) amplification (mean 19 copies) and none of 78 patients with negative PD-L1 expression had CD274 amp (P = .05). In contrast, 4.2% of the patients with high PD-L1 expressing UBC had FGFR3 alterations while 26.6% of those with negative PD-L1 expression did (p=0.02).
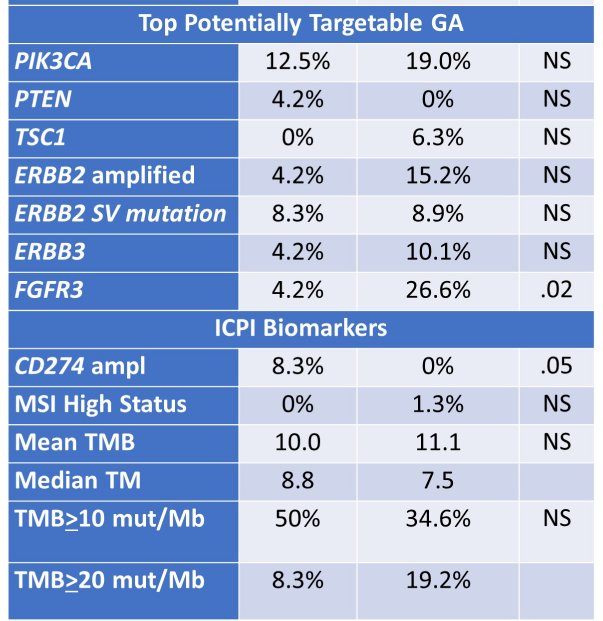
While not statistically significant, a number of currently “untargetable” genomic alterations were more frequent in the PD-L1H UBC including TP53, TERT, and RB1. MTAP loss, a potential target for PRMT5 and MTA2 inhibitors, were three times more frequent in the PD-L1 negative UBC (25.3% vs 8.3%) though this was not significant. Similarly, ERBB2 amplification and ERBB3 and PIK3CA short variant (SV) genomic alterations were more frequent in the PD-L1 negative groups though differences did not reach significance. Other ICPI-associated potential biomarkers, including MSI status, TMB level and GA in PBRM1, STK11 and MDM2 were not significantly different in the groups.
Among those patients where a mutational signature could be determined, 10/12 (83%) of patients with high PD-L1 expression and 21/29 (72%) of patients with PD-L1 negative UBC featured APOBEC signature. Interestingly, 2 PD-L1 negative tumors featured an MMR signature and 6 PD-L1 negative tumors featured no dominant signature.
Thus, the authors conclude that the genomic profiles of high and negative PD-L1 expressing tumors differ, with a greater proportion of potentially targeted alterations among those with negative PD-L1 expression, including FGFR3, ERBB2, ERBB3, and PIK3CA. PD
Presented by: Petros Grivas MD, Ph.D., University of Washington and Fred Hutchinson Cancer Research Center, Seattle, WA
