He first showed a timeline of the history of research on what became known as the Von Hippel-Lindau (VHL) genes. Retinal angiomatosis, a disorder of inherited alterations in the VHL gene were first described at the end of the 19th century, and the term VHL disease was coined in the 1930s. Work in the late 1980s and early 1990s from W. Marson Linehan and others led to an understanding of the genetic basis of VHL disease by mapping the location of the VHL locus on chromosome 3p. Indeed, the genomic hallmark of ccRCC is alterations of chromosome 3p. Subsequently, studies by Bill Kaelin and others linked the VHL disease to HIF (hypoxia-inducible factor) signaling and RCC survival. These have led to many therapies targeting the VHL-HIF-VEGF signaling pathways in RCC.
RCC is a disease dominated by alterations in tumor suppressors causing loss of function. In addition to alteration in VHL, mutations in other genes on chromosome 3p such as PBRM1, SETD2, and BAP1 have been identified recurrently in various RCC subtypes. Other notable mutations in RCC include alterations in the PI3K/mTOR signaling pathway, although these tend to be more subclonal, which may explain the limited efficacy of mTOR inhibitors in RCC.
More recent multiregional sampling studies have further elucidated key characteristics of ccRCC pathogenesis. Specifically, it is thought that the first early event is loss of one allele of chromosome 3p, followed years later by VHL inactivation and often an amplification of chromosome 5q. Subsequently, after the acquisition of other clonal and subclonal drivers, ccRCC ultimately develops. This is illustrated in the schematic below. Additionally, multiple molecular subtypes driving primary tumor growth have been identified, including a VHL mono-driver state, BAP1 driven tumors, and multiple contexts of PBRM1 mutation (Figure below from 10.1016/j.cell.2018.03.043).1
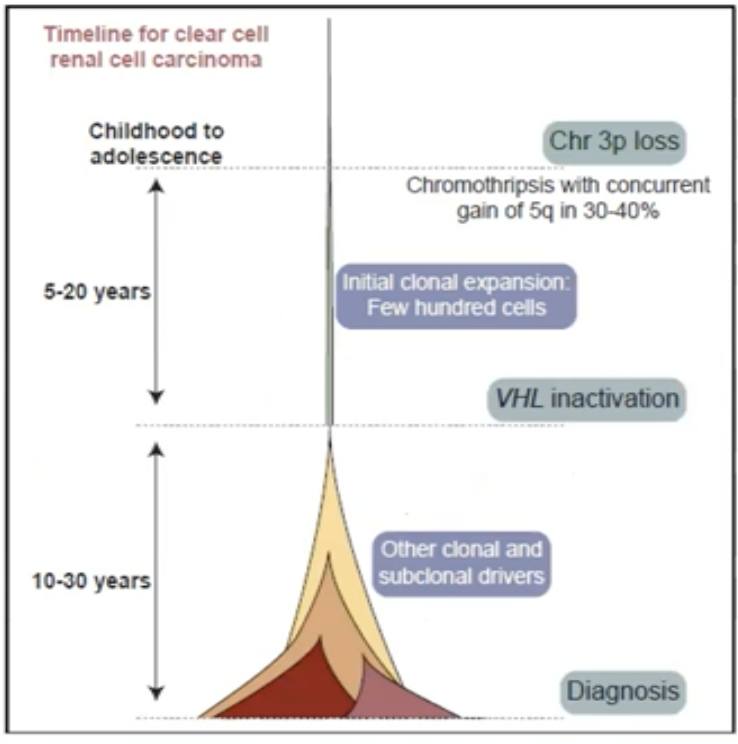
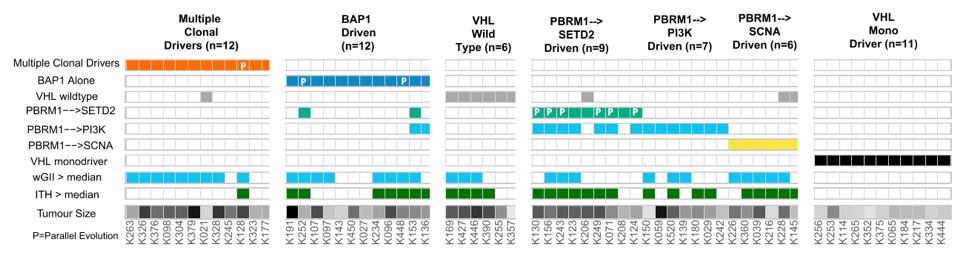
The metastatic process selects for certain mutation and copy number events also, including loss of chromosome 9p, which harbors the CDK2NA gene. Other genomic drivers seem to be prognostic. For example, alterations in PBRM1 are present in 40-50% of all tumors and are associated with better overall prognosis. However, mutations in SETD2 and BAP1 are associated with worse prognosis. Mutations in BAP1 especially are associated with higher-grade disease whereas mutations in SETD2 drive subclonal diversity and metastatic potential.
Finally, Dr. Hakimi focused on the immune milieu of ccRCC. He first highlighted work forthcoming from his group using single cell profiling of ccRCC that identifies significant infiltration of CD8 T cells and tumor-associated macrophages in these tumors. He then highlighted recent work showing that different expression patterns (angiogenesis, immune cold, T-effector, etc in the tumor microenvironment define response.2 Additionally, certain genomic drivers are linked to unique tumor microenvironment characteristics. For example, PBRM1 mutations are associated with a more angiogenesis expression program in the microenvironment, and a more myeloid microenvironment is associated with CDKN2A/B loss and TP53 alterations.
Presented by: A. Ari Hakimi, MD, Urologic Surgeon Memorial Sloan Kettering Cancer Center
References:
1. Turajlic S, Xu H, Litchfield K, et al., Deterministic Evolutionary Trajectories Influence Primary Tumor Growth: TRACERx Renal. Cell, 2018; 173(3);:595-610
2. Huseni M D, Atkins M, et al.,Clinical activity and molecular correlates of response to atezolizumab alone or in combination with bevacizumab versus sunitinib in renal cell carcinoma. Nature Medicine, 2018; 24:749–757

